Functional normalization
Motivation
Our workflow uses the functional normalization approach16, which exploits internal control probes designed to detect technical variations without assaying biological differences.
It has been shown to perform favourably when compared to other approaches17. Using the internal control probes avoids the problems associated with global normalization methods, where biological variation can be mistaken for a technical effect and removed. This is especially important in studies where groups are expected to have differential methylation signatures, such as multiple tissue studies18.
Conversations on the best approaches for normalization in DNAm data pipelines are ongoing19.
Principal Components
The default of selecting only two principal components is often too low for this type of data. Often you will see a drop-off in proportion of variance explained after a certain number of principal components, and this can indicate an efficient selection.
var_explained %>% ggplot(aes(x=PC, y=var_explained)) +
geom_line() +
geom_point(color='grey5', fill='#6DACBC', shape=21, size=3) +
scale_x_continuous(breaks=1:ncol(pca$x)) +
xlab("Principal Component") +
ylab("Proportion of variance explained") +
theme_bw()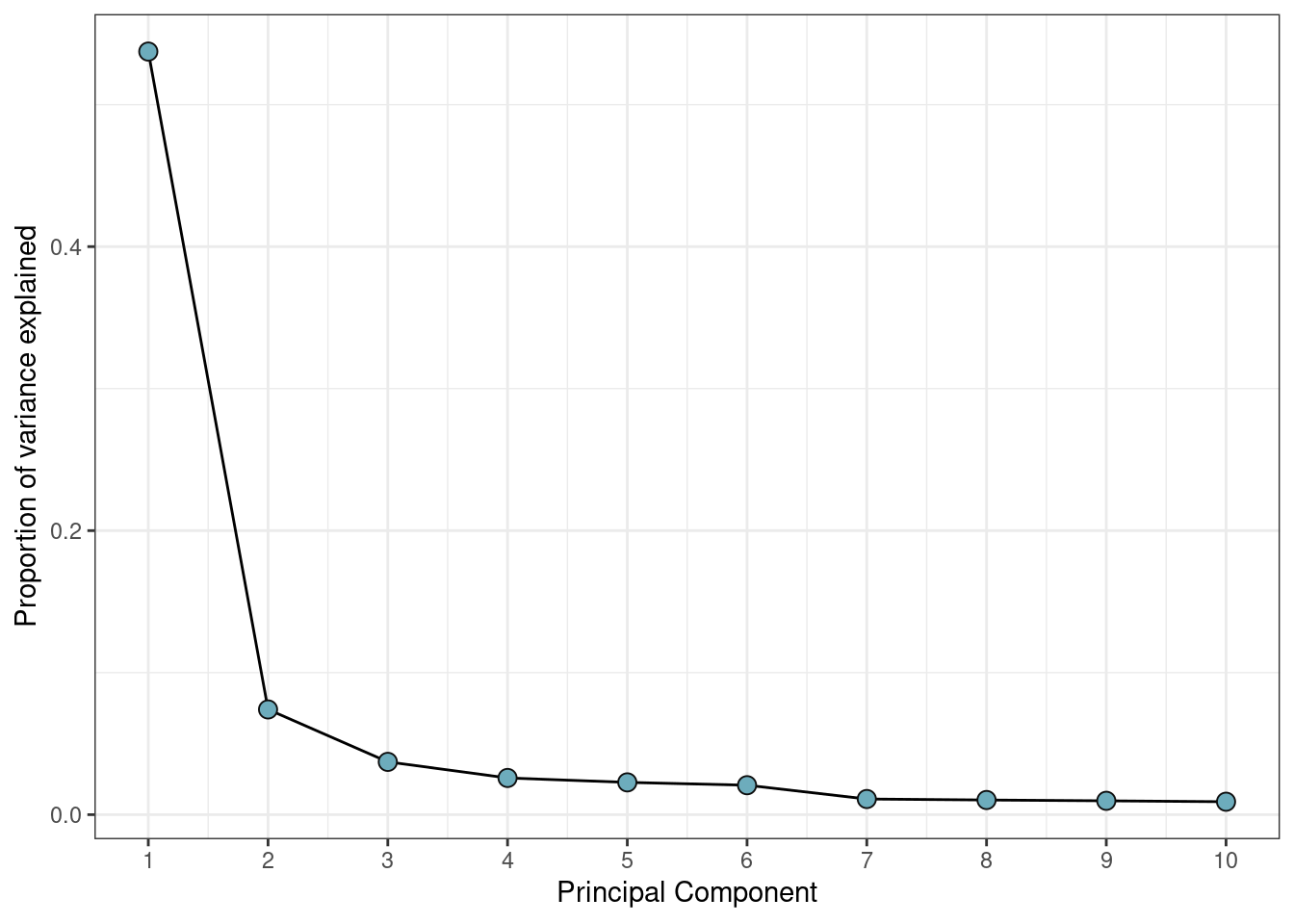
Running Normalization
In order to run normalization the annotation of the
RGset must be updated for EPIC arrays.
By default, functional normalization returns normalized copy number
data making the returned GenomicRatioSet twice the size
necessary when only beta-values or M-values are required. Therefore, we
developed preprocessFunnorm.DNAmArray(), which adds an
option not to return these to the preprocessFunnorm()
function from minfi.
You can now extract the normalized beta values from this object. It can also be advisable to round the beta values to 4 significant figures, which both reduces the data size and provides a realistic measure of the available precision.
If required, these can be transformed to M-values using minfi’s
getM() which applies a logit2-transformation to the
beta-values.